ENS - PSL |Laboratoire PASTEUR

Started on Oct. 16, 2023
In pharmacology, a drug is a chemical substance which, when administered to a living organism, produces a biological effect. There are two main families of drugs: biological molecules (extracted or produced by a biological organism) and "small molecules" (synthetic). It is to the latter category, which includes molecules with a size of between ten and a hundred atoms, that the majority of drugs placed on the market each year belong, even if the share of other drug families increases over time. Most often, a drug targets a protein and seeks to inhibit its activity.
Proteins are molecules made up of one or more chains of amino acids, which perform a variety of functions in the human body, including catalyzing chemical reactions, transporting molecules, and transmitting signals between cells. Proteins are essential to the normal functioning of the human body, but when they are altered or dysfunctional, this can contribute to the development of diseases.
The first crucial step in drug discovery is the identification of hits. This involves screening large libraries of compounds to identify molecules with interesting biological activity against a specific target. Virtual screening has become a popular method for this step because it enables millions of compounds to be screened using computer simulations, rather than performing physical tests on each compound, as is done in traditional high-throughput screening. By using machine learning models to predict a molecule's ability to interact with a specific target, researchers can identify hits more quickly and reduce the costs associated with drug discovery. Therefore, an area of research in the pharmaceutical industry is to develop powerful models to predict the interaction of molecules with biological targets, such as proteins. These models are also very useful for the subsequent drug discovery process to predict the activity of different drug candidates.
Can you predict molecular properties to discover new drugs ?
The challenge proposed here is a supervised regression problem, the objective being to predict the inhibitory capacity of molecules on a protein. The pIC50 is a measure of the inhibitory capacity, it is calculated from the half-inhibition concentration (IC50) of the compound, which is the concentration of the compound necessary to inhibit 50% of the target activity. The higher the pIC50 of a molecule, the higher its inhibitory capacity. Generally, a molecule is considered active if its pIC50 is greater than 8. We are therefore in a context of drug discovery, where we have molecules whose activity on a protein is known, with which we will build a QSAR (quantitative structure-activity relationship) model to predict the activity of new molecules.
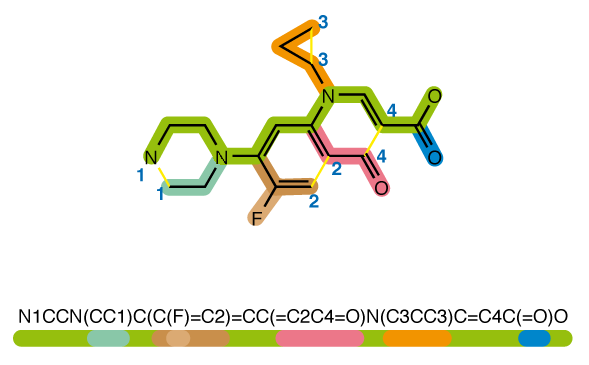
The molecules are stored in the SMILES (simplified molecular-input line-entry system) format, which allows a representation in the form of a string. It is possible to work directly with this format but it is also possible to convert SMILES into molecular graphs or molecular fingerprints. Fingerprints are binary vectors that capture the presence or absence of specific chemical features, such as functional groups, atom types, bonds, or molecular properties. It is the most common way to encode the chemical information contained in a molecule and that is the approach used in the introduction notebook to get the baseline score.

You have at your disposal 4 csv files:
X_train.csv : contains the unique identifiers of 4400 encoded molecules and the associated SMILES
y_train.csv : contains the unique identifiers of 4400 molecules and the associated pIC50 value
X_test.csv : contains the unique identifiers of 2934 molecules and the associated SMILES
random_submission_example.csv : contains an example of submission in the right format
supplementary_files : contains a notebook to introduce the challenge and replicate the baseline score.
The objective is to predict the pIC50 values of the molecules contained in the X_test.csv file and submit them on the platform.
id : unique identifiers of each molecule,
smiles : molecules in SMILES format,
y : values of the pIC50.
All data are extracted from the ChEMBL database.
The metric used is the MAE (Mean Absolute Error) defined by :

where n is the number of elements, Yi the actual value and Ŷi the predicted value.
A baseline score was obtained by converting the SMILES to Morgan fingerprints and then training a Random Forest algorithm with the default parameters from scikit-learn. You can reproduce this score by using the introduction notebook associated with the challenge.
You can find a notebook to reproduce the baseline score in the supplementary files.
Files are accessible when logged in and registered to the challenge
